A groundbreaking study by Indian scientists has overturned a 50-year-old textbook model of bacterial gene regulation, challenging the long-accepted “σ (sigma) cycle” theory and opening new pathways for understanding bacterial gene regulation and its evolution.
The research, conducted by scientists at the Bose Institute and Rutgers University, reveals that the sigma cycle – long believed to be universal in bacterial gene transcription – does not apply to all bacteria.
The findings were published in the Proceedings of the National Academy of Sciences (PNAS).
Study Overturns Central Model of Bacterial Gene Transcription
For nearly five decades, biology textbooks have described how bacteria turn their genes on through the “σ (sigma) cycle.” In this widely accepted model, sigma factors bind to RNA polymerase to initiate transcription and then dissociate to allow elongation.
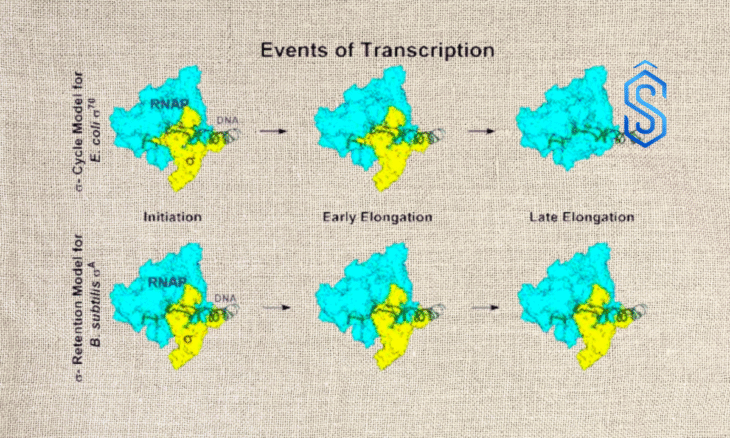
The model was largely built on observations from the bacterial strain Escherichia coli σ70.
However, the new study demonstrates that this cycle is not universal in bacterial gene regulation.
Researchers reported that, contrary to decades of scientific belief, the principal transcription initiation factor in Bacillus subtilis—σA—and a modified version of the Escherichia coli σ70 factor remain bound to RNA polymerase throughout transcription, instead of being released after initiation.
“Our work shows that in Bacillus subtilis, the σA factor stays attached to RNA polymerase all the way through the transcription process,” said Jayanta Mukhopadhyay, corresponding author from the Bose Institute.
“This fundamentally changes how we think about bacterial transcription and gene regulation.”
Also Read: Recombinant Cells and Sensors Facility Launched at BRIC-RGCB with INR 60 Cr DBT Investment
Real-Time Observation Reveals New Bacterial Gene Mechanism
To investigate bacterial gene behaviour, researchers used a combination of modern techniques including biochemical assays, chromatin immunoprecipitation, and fluorescence-based imaging.
These tools enabled them to observe sigma factor behaviour in real time.
The team found that Bacillus subtilis σA and an E. coli σ70 variant lacking a part called 1.1 remain stably associated with transcription complexes. This is in stark contrast to full-length E. coli σ70, which is released stochastically during elongation.
“These findings provide compelling evidence that the long-accepted σ cycle does not apply to all bacteria,” added co-author Aniruddha Tewari of the Bose Institute. “It opens new avenues for understanding bacterial gene regulation and its evolution.”
The discovery significantly reshapes current understanding of bacterial gene transcription mechanisms and suggests that bacterial gene regulation may vary more widely across species than previously believed.
Also Read: Aqueous Zinc-Ion Batteries for Grid Storage: CeNS Scientists Develop Advanced 1T-MoS₂ Cathode
Bacterial Gene Regulation: Implications for Antibiotics, Bioengineering and Infection Control
The findings have broad implications for microbiology and biomedical research. A deeper understanding of bacterial gene regulation could influence how researchers approach bacterial physiology, stress response, and antibiotic development.
By redefining how bacterial gene transcription operates in certain species, the study may contribute to designing better antibiotics or regulatory inhibitors that block infection mechanisms.
Additionally, it may support the development of engineered microorganisms capable of efficiently producing biofuels, biodegradable plastics, or therapeutic compounds.
As scientists revisit long-standing assumptions about bacterial gene transcription, this study marks a significant milestone in molecular biology and could reshape research strategies in microbiology and biotechnology.
Authors and Institutions
The study was authored by Aniruddha Tewary, Shreya Sengupta, Soumya Mukherjee, and Nilanjana Hazra from the Bose Institute, and Y.W.E., R.H.E., Yon W. Ebright, Richard H. Ebright, and Jayanta Mukhopadhyay from Rutgers University, USA.